 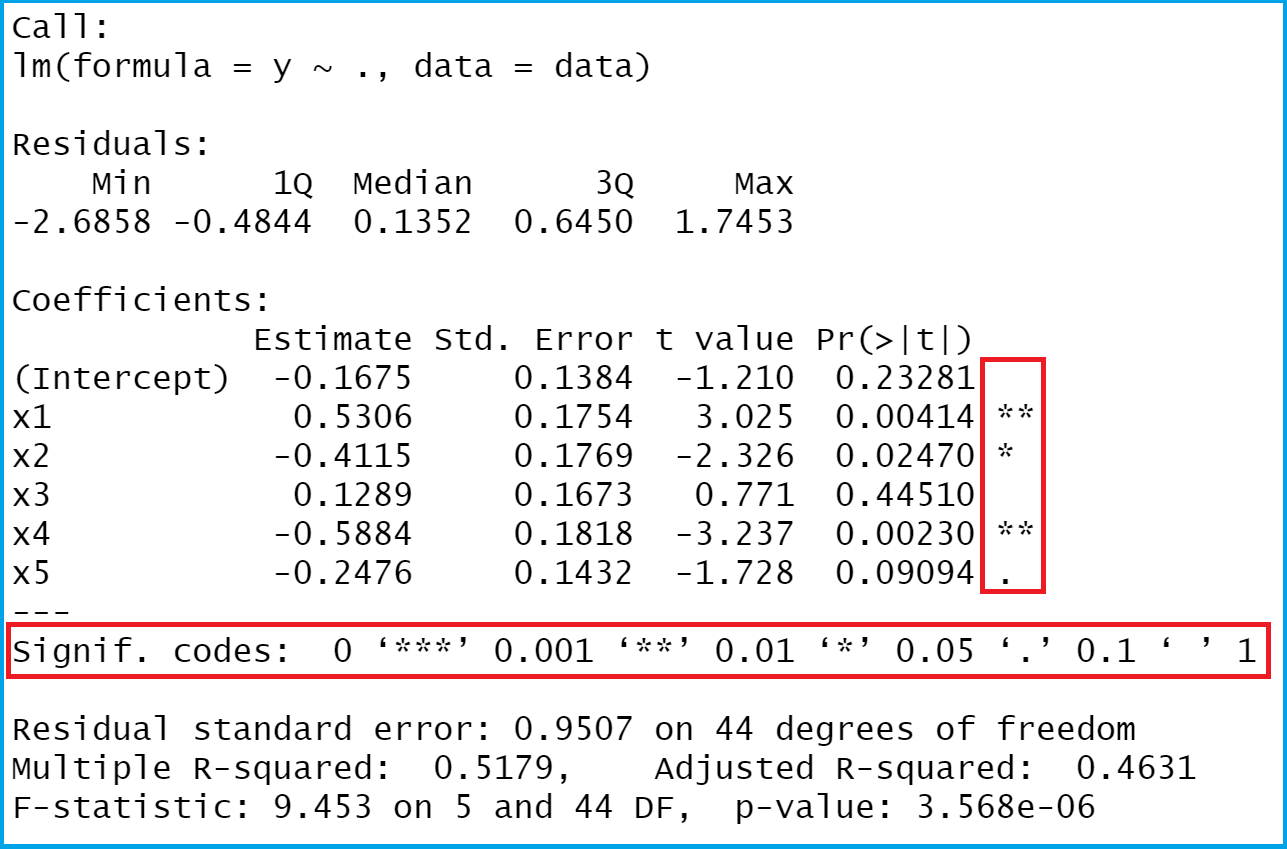
In this example, we can observe an ‘elbow’ around PC9-10, suggesting that the majority of true signal is captured in the first 10 PCs. While still available in Seurat (see previous vignette), this is a slow and computationally expensive procedure, and we is no longer routinely used in single cell analysis.Īn alternative heuristic method generates an ‘Elbow plot’: a ranking of principle components based on the percentage of variance explained by each one ( ElbowPlot() function). In Macosko et al, we implemented a resampling test inspired by the JackStraw procedure. However, how many components should we choose to include? 10? 20? 100? 
The top principal components therefore represent a robust compression of the dataset. To overcome the extensive technical noise in any single feature for scRNA-seq data, Seurat clusters cells based on their PCA scores, with each PC essentially representing a ‘metafeature’ that combines information across a correlated feature set. For example, in Seurat v5, the count matrix is stored in pbmc]$counts.ĭetermine the ‘dimensionality’ of the dataset For more information, check out our, or our GitHub Wiki. The object serves as a container that contains both data (like the count matrix) and analysis (like PCA, or clustering results) for a single-cell dataset. We next use the count matrix to create a Seurat object. Note that more recent versions of cellranger now also output using the h5 file format, which can be read in using the Read10X_h5() function in Seurat. The values in this matrix represent the number of molecules for each feature (i.e. gene row) that are detected in each cell (column). The Read10X() function reads in the output of the cellranger pipeline from 10X, returning a unique molecular identified (UMI) count matrix. There are 2,700 single cells that were sequenced on the Illumina NextSeq 500. Demultiplexing with hashtag oligos (HTOs)įor this tutorial, we will be analyzing the a dataset of Peripheral Blood Mononuclear Cells (PBMC) freely available from 10X Genomics.Analysis of spatial datasets (Sequencing-based).Analysis of spatial datasets (Imaging-based).Map COVID PBMC datasets to a healthy reference.Sketch integration using a 1 million cell dataset from Parse Biosciences.Integrating scRNA-seq and scATAC-seq data.Dictionary Learning for cross-modality integration. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
